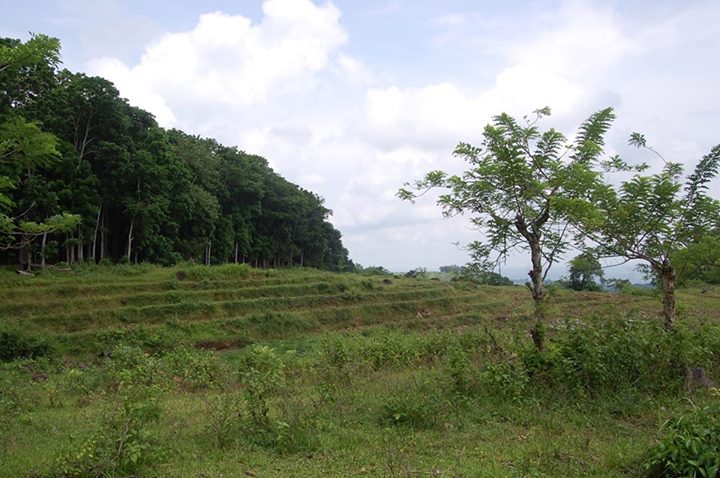
Ph.D. ExperienceForest Fragmentation in the Philippines
For my Ph.D., my focus was on forest fragmentation, which threatens the survival of many species due to reduced habitat and increased restrictions on dispersal and gene flow. Specifically, I was interested in quantifying the influence of edges on remaining forest stands. Research on changes at forest edges has revealed that edge influence can lead to the further degradation of forest fragments. Furthermore, edge-affected areas are less able to support viable populations of edge-sensitive species, effectively further reducing the area of suitable habitat for many species. My research brought me to the Philippines, a world leader in biodiversity, endemic species, and unfortunately, deforestation. I went to the Philippines twice, in May 2013 and 2014, the latter trip funded by National Geographic. The Phylogeography of Southeast Asian Butterflies During my time in Dr. Lohman's lab, I also did a lot of bench work generating molecular data for a phylogeography project concerning butterflies in Southeast Asia. Butterflies are highly diverse (approximately every 1 out of every 12 animal species is a butterfly) and Southeast Asia is not only a biodiversity hotspot, but it's also extremely under researched. Master's ExperienceMy Master's Project
My Master’s project investigated the post-transcriptional regulation of the rate-limiting, nitrogen assimilating gene nitrate reductase (NR, encoded by nia) in the model marine diatom Thalassiosira pseudonana. The assimilation of nitrogen is a critical control of marine phytoplankton growth, and is a highly regulated process in marine diatoms. To study these multiple levels of control, I created transgenic lines of T. pseudonana containing DNA constructs that drove the nitrate-inducible expression of the reporter EGFP (enhanced GFP). This project required mastering many different molecular techniques, such as molecular cloning, developing primers, Sanger sequencing, RNA extractions, cDNA synthesis, standard PCR, real-time PCR, genetic transformation via particle bombardment, culturing bacteria and marine diatoms, and creating alternative approaches to bypass experimental roadblocks. I presented this work in a talk at the 51st Annual Northeast Algal Society Symposium. Following this conference, my project concluded with a Master’s defense in the style of a departmental seminar. Both experiences were close to nerve-shattering, but in the end unforgettable life and professional experiences. |
|
Undergraduate Experience
I was involved with research projects throughout my undergraduate career at Clark University. During the fall semester of my first year, I took a seminar course in which we collected tissue samples from fungal fruiting bodies and associated parasitic plants in order to determine any possible phylogenetic relationship. Through the amplification of ITS sequences from extracted DNA, we found genera-specific plant-fungal associations.
My senior year I developed, led, and carried out an original project to test whether juvenile body size of the threespine stickleback (a model for evolution and adaptive radiation) plays an important role in competition for limited food sources. Our data provided evidence that suggested larger juveniles were better competitors for food than their smaller counterparts. Because larger body size is critical in the overwintering of fish, traditionally a period of high mortality, this project could have significant implications for threespine stickleback life history traits. Both of these projects are examples of my continued desire to broaden my horizons within the area of ecology, evolution, and behavior, in addition to strengthening my writing and communication skills in areas of biology that I am unfamiliar with.
My first project in the Robertson Lab involved collecting DNA sequences from online databases and assembling gene trees for a Ph.D. student. The following semester, I was involved with cloning untranslated regions (UTRs) and designing primers, which supported the Master’s project of my lab partner. The following summer, I was awarded a fellowship to begin what would become my Master’s thesis project, which I completed May 2012. This work was conducted over two summers and four semesters (two as an undergraduate and two as an accelerated Master’s student).
My senior year I developed, led, and carried out an original project to test whether juvenile body size of the threespine stickleback (a model for evolution and adaptive radiation) plays an important role in competition for limited food sources. Our data provided evidence that suggested larger juveniles were better competitors for food than their smaller counterparts. Because larger body size is critical in the overwintering of fish, traditionally a period of high mortality, this project could have significant implications for threespine stickleback life history traits. Both of these projects are examples of my continued desire to broaden my horizons within the area of ecology, evolution, and behavior, in addition to strengthening my writing and communication skills in areas of biology that I am unfamiliar with.
My first project in the Robertson Lab involved collecting DNA sequences from online databases and assembling gene trees for a Ph.D. student. The following semester, I was involved with cloning untranslated regions (UTRs) and designing primers, which supported the Master’s project of my lab partner. The following summer, I was awarded a fellowship to begin what would become my Master’s thesis project, which I completed May 2012. This work was conducted over two summers and four semesters (two as an undergraduate and two as an accelerated Master’s student).